Machine learning that uses seismocardiogram (SCG) signals is limited by lack of data
The SCG signal, which measures chest wall vibrations caused by heartbeats, is increasingly used in cardiovascular health assessments. It provides clinically relevant information related to aortic opening (AO) and aortic closing (AC) events, which are important features for monitoring heart failure, detecting the acute physiological effects of non-invasive neuromodulation and stressors, and estimating a variety of hemodynamic variables (e.g., blood pressure, stroke volume).
However, the lack of SCG data available for training is limiting machine learning approaches that leverage SCG signals. This is especially the case for deep learning where large amounts of data (including diverse datasets) are required to achieve broad and reliable generalized results without overfitting. In addition, safety concerns related to using human participants as well as difficulty replicating the high-noise environments found in real-world applications creates a lack of access to ground truth labels, which are used to compare and evaluate model results.
Phantom generator hardware and deep generative models create synthetic data that replicates human body signals
By combining phantom generator hardware and deep generative models to create synthetic data, this system provides expanded datasets that are then naturally mixed with environmental noise. It addresses dataset scarcity while enabling safe and secure data collection in environments that are unsafe for human experiments but relevant to real-world scenarios (e.g., surgical research, patients in critical conditions, vulnerable populations).
The system uses hardware to replicate human body signals, including those measured by both SCG and electrocardiogram (ECG), and a deep generative model to create synthetic SCG beats with clinically relevant SCG features. This synthetic data generation system introduces diversity and control over signal parameters and enables bio-signal data collection without human participants.
The hardware system was validated using human SCG and ECG signals and strong correlations between features of the generated SCG beats and the desired features fed into the model. The deep generative model produces physiologically diverse, realistic SCG signals and provides precise control over AO and AC features. This uniquely enables dataset augmentation for SCG processing and machine learning to overcome data scarcity.
- More realistic: The transformer-based generative model synthesizes SCG beats that are more similar to real human SCG beats than porcine SCG beats and are less similar to non-SCG signals. The signals replicated using the phantom generator showed correlations to real human signals in both the time domain (greater than 90%) and the frequency domain (greater than 98%).
- Controllable: Clinically relevant feature inputs act as control knobs for the AO and AC aspects of the generated SCG beat.
- Diversity enabling: The model enables randomization of beat morphology through a random identification token. The token introduces diversity in the morphology of the SCG signals while maintaining control over physiological variation—a desirable feature for dataset augmentation.
- Generalizable: For the deep generative model, the distance between the training dataset and the test dataset is very close to the distance between the synthetic dataset and the training dataset, showing that the model generalizes reasonably.
- Portable: The hardware is a portable, USB-C powered system based on a Raspberry Pi® computer that can replicate any SCG and ECG signal inputs with minimal error in acceleration and voltage.
- Versatile: Both the generative hardware and the deep generative model are expandable to other bio-signals, such as photoplethysmogram (PPG), phonocardiogram (PCG), electroencephalogram (EEG), respirogram, etc.
Raspberry Pi is a trademark of Raspberry Pi Ltd.
- Data augmentation for healthcare artificial intelligence training databases:
- Cardiovascular signal processing
- Other pre-trained models for pre-ejection period (PEP) and left ventricular ejection time (LVET) estimation or disease classification
- Other bio-signal training sets
- Denoising algorithms, quality indexing, and feature discovery:
- Complex deep learning models for health assessment based on bio-signals
- Data collection in very noisy and unsafe environments (e.g., for military health assessments, in high-vibration environments)
- Training modules to remove motion noise artifacts from bio-signals
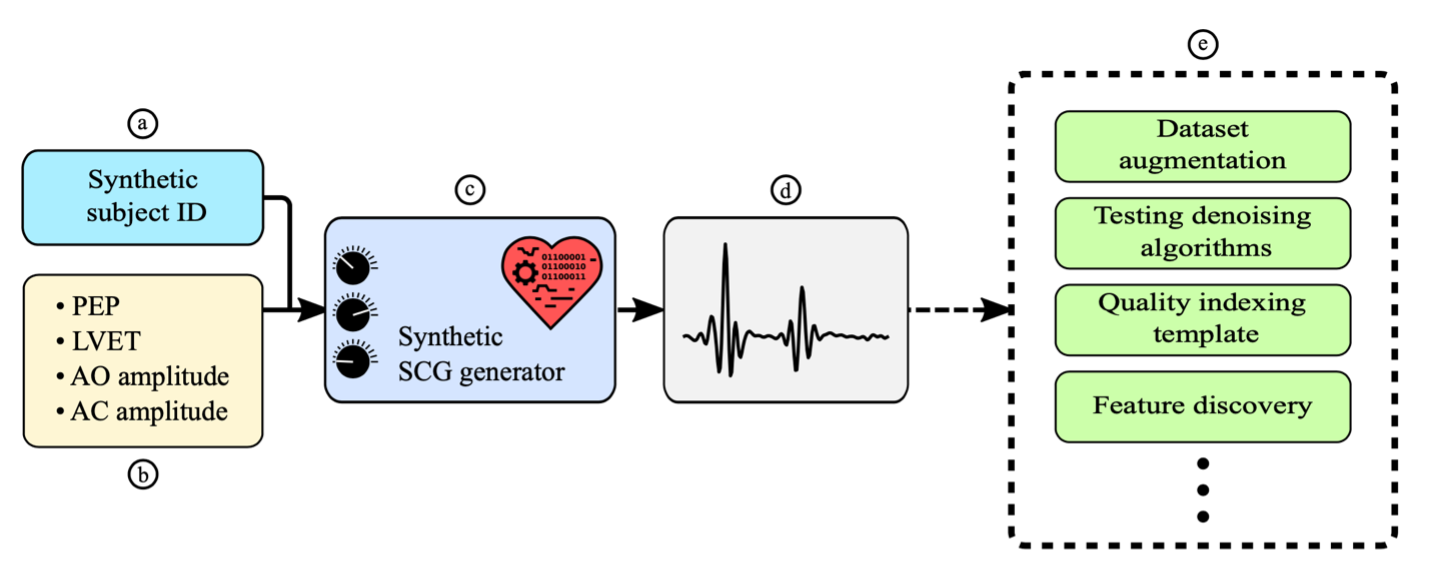
Overview of the synthetic SCG generator model. (a) A participant identification (ID) is fed as input, which controls the morphology of the generated beat. (b) The model receives SCG features as inputs and then outputs an SCG beat with those features. (c, d) This model can be used to generate sequences with controlled features. € This enables tasks such as dataset augmentation and template generation. It also sets the stage for simulating noisy SCG and testing denoising algorithms.
